Introduction
At the Gilleard lab meeting on December 16, Dave brought to my attention an unfortunate oversight in my BLAST visualization tool Kablammo, which I introduced when I launched it in July. My error was in ignoring strandedness–when I wrote Kablammo, I put insufficient thought into the importance of such information. I simply wished to visualize alignments between the query and subject sequences, and in so doing blundered forth like a birdbrained buffoon.
Two issues arise when one ignores BLAST strandedness. Allow me to illustrate the first.
How to visualize stranded hits
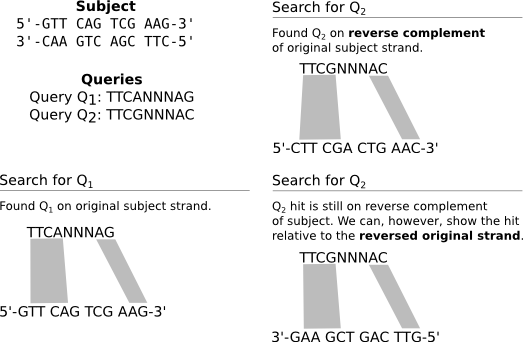
Observe that you have a single subject sequence (presented both in its forward and reverse-complement forms) against which BLAST searches, as well as two distinct queries. Our goal is to find alignments between each query and the subject, ignoring the three-N run in the middle of the queries. To appreciate this example, you must understand that BLAST will report all hits relative to the original subject sequence, even when the hit lies on the subject’s reverse complement. Three scenarios result:
-
We search for Q1, finding it in the original subject. We can simply display the subject sequence from 5’ to 3’, and all is well.
-
We search for Q2, finding it in the reverse complement of the subject. In this case, we can display the reverse complement of the subject from 5’ to 3’, and all is well.
-
Suppose we wish to avoid displaying the subject’s reverse complement, and instead display the hit for Q2 relative to the original subject sequence. This is what BLAST does, and for good reason–it simplifies the display of results, for it means that all sequence coordinates reported are relative to the original subject sequence regardless of strand. In this case, we can show the Q2 hit relative to the reversed subject, meaning we render the subject from 3’ to 5’.
Scenario 3 is where Kablammo previously faltered–in hits to the reverse complement of the subject, it would render the subject from 5’ to 3’, rather than 3’ to 5’. In the scenario above, you can see this would result in an intersection between the two trapezoids indicating hits. How does this look in practice?
How to suck at visualizing things, part I
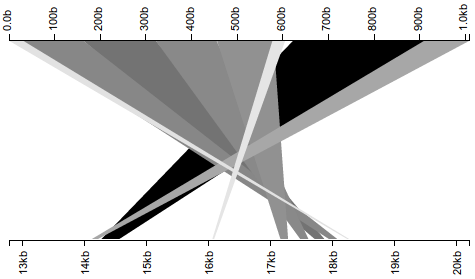
Observe that the subject query (bottom axis) is rendered from 5’ to 3’ (meaning that sequence coordinates increase from left to right). This appears to be a clear case of misassembly–though almost the entirety of the query sequence (top axis) is present in the subject, the subject’s assembly seems to be out-of-order. When I presented this figure, however, Dave noted that the seeming misassembly appears precisely inverted–i.e., if I inverted my subject axis such that it ran 3’ to 5’ (meaning that sequence coordinates would increase from right to left), the sequence would instead appear well-assembled. What happens when I perform this inversion?
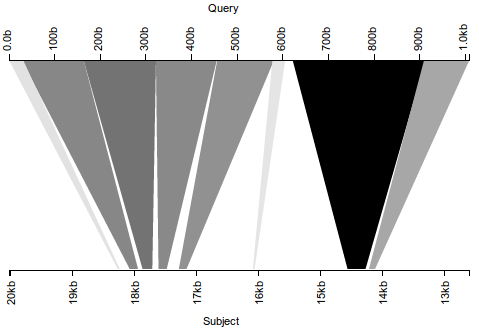
Note that, with the subject axis inverted, all fragments are ordered properly. (Gaps in the alignment correspond to introns, which we expect–the original query sequence consisted solely of a gene’s exons.) A seemingly minor change to the visualization thus leads to a completely different conclusion–whereas before, this was a clear instance of misassembly, it is now a well-assembled gene.
How to suck at visualizing things, part II
This, alas, does not bring us to the end of our journey. Once I began inspecting Kablammo’s visualizations, I stumbled upon a second problem. Though Kablammo was now adjusting for strand, reversing the subject axis when hits lay on the subject’s reverse complement, I was ignoring the possibility that one may have hits to both the original subject and its reverse complement. Hits to both strands of a given subject are relatively rare, but do nonetheless occur. In such cases, I rendered both on the same plot, leading to a visualization such as this:
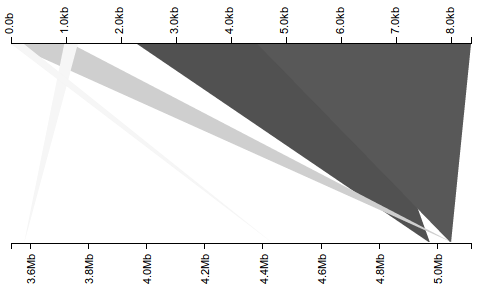
Here, we again see what appears to be a misassembly, with portions of the query sequence scattered throughout the subject, and most notably, almost 2 kb of overlap between the two dark triangles corresponding to the latter two-thirds of the query sequence. If, however, we properly separate the hits into sets corresponding to the original subject and its reverse complement, we see an altogether different result.
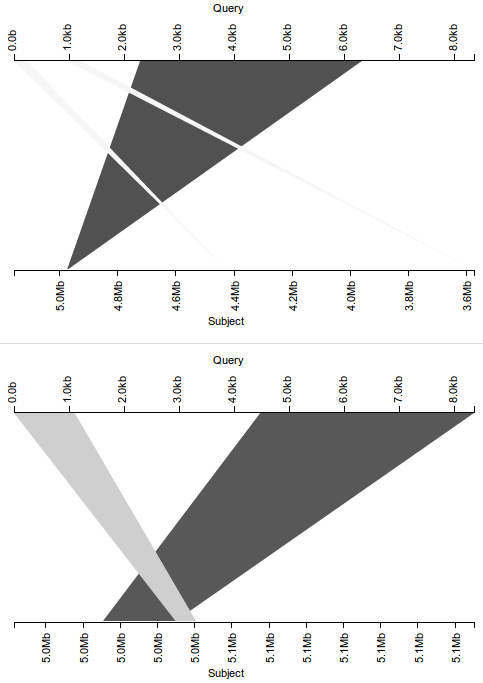
With the five hits now separated by strandedness across two plots, we see the two dark triangles actually correspond to hits on different strands, eliminating the overlap so suggestive of misassembly. We still see evidence of misassembly–consider the lower plot corresponding to the non-reverse-complement subject, showing that the first kilobase of the query appears out-of-order in the subject–but the problem seems substantially less dire than before. Again, a small adjustment to the visualization leads to a substantially different conclusion.
Conclusion
So, what have I learned on my journey?
-
Hack responsibly. When I first wrote Kablammo, I did it as a quick hack that I gradually expanded into a production application. In my first pass, I was so eager to get something working that I didn’t stop to think carefully about strandedness, instead concerning myself only with how to produce a picture with minimal effort.
-
Don’t trust your visualizations. I trusted Kablammo implicitly because it produced reasonable-looking results, and so I gradually grew accustomed to its results such that I never returned to verify their veracity. Only when Dave critically examined my plots did my error come to my attention; without Dave’s fresh perspective, I likely would have gone on producing erroneous plots for some time to come.
-
Fixing fundamental issues in software really sucks. Though the number of lines of code I’ve changed to fix this problem is minimal, figuring out the precise nature of the problem, how best to resolve it, and then verifying that the changed system correctly renders BLAST hits has taken me nine or ten hours across the past three days. Writing code for sexy new features is much more fun.
With all the above said, I now have more faith in Kablammo’s fidelity. If that isn’t worth hours hunched over my keyboard, I don’t know what is.