For the last few weeks, I’ve been hacking sporadically on Kablammo, a web-based tool for visualizing BLAST results. For each alignment produced by BLAST, Kablammo will illustrate exactly which portions of the query sequence (i.e., the sequence you gave BLAST) map to which portions of the subject sequence (i.e., the sequence BLAST found in its database that aligns to your input). This can be of considerable value in visualizing genome misassembly.
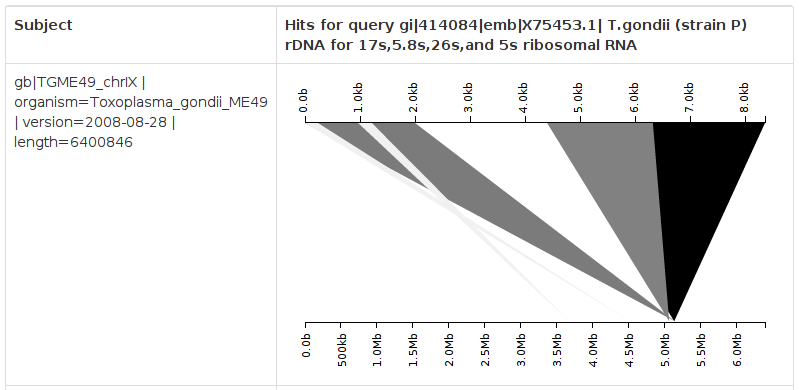
In this case, the query sequence, seen on the top axis, is the Toxoplasma gondii rRNA cassette, which was drawn from GenBank and corresponds to the correct sequence in vivo. The subject sequence, seen on the bottom axis, is the Toxoplasma reference genome. Each alignment produced by BLAST is coloured according to its E value, with higher-quality hits coloured darker. We see clearly that, for this BLAST hit, five separate portions of the query sequence align to five portions of the Toxoplasma chromosome IX reference assembly, suggesting that the reference is erroneously assembled. Furthermore, the region from approximately 2.0 kb to 4.5 kb is not present within the reference assembly, lending further credence to our conclusion.
Now, you might be wondering, “Why should I use Kablammo?” Well, by golly, I’m glad you asked!
-
Kablammo is entirely web-based. You need not muck around with anything on your local machine–just open it in a respectable web browser such as Firefox or Chrome, then start visualizin’.
-
Kablammo’s graphs are rendered as SVGs. This means you can scale them to any resolution for use digitally or in print. Furthermore, you can save the SVG graphs using your web browser, then open them in a vector graphics editor such as Inkscape or Adobe Illustrator to manipulate them as you please.
-
Kablammo is open-source.
-
Kablammo is ridiculously easy to run on your own server. It doesn’t rely on any server-side code, for it is written entirely in JavaScript and executes solely within the browser. This means that, to host your own copy, you need only serve it as a set of static files. Deployment can be as easy as downloading Kablammo from GitHub, then running
python3 -m http.serverfrom its directory.
I humbly hope that Kablammo may be of use to someone else floating in the ether.